Every real dataset has holes. A column full of blanks, a timestamp that simply never arrived, a field the respondent left empty — on the surface they all look the same. In practice they are not. The reason a value is missing is as important as the value itself, and conflating different reasons leads to models that are quietly, confidently wrong. A patient cohort where sicker people drop out of a trial is a different problem than a sensor that randomly powers off. Imputing both the same way is the kind of mistake that passes cross-validation and fails in production.
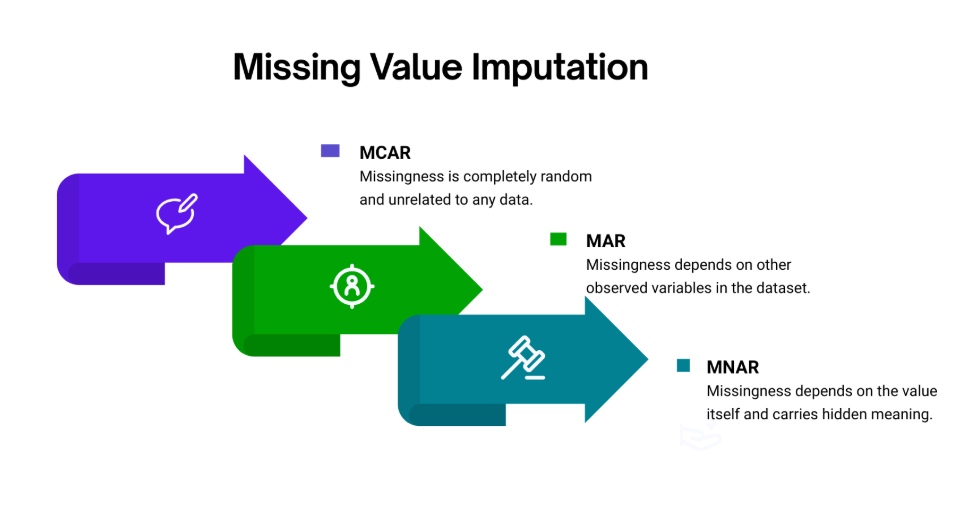
The field has three named mechanisms for why values go missing. MCAR — missing completely at random — is the benign case: the absence of a data point has nothing to do with anything measured or unmeasured. MAR — missing at random — is more subtle: the probability of missingness depends on other observed columns, but not on the value that is actually missing. MNAR — missing not at random — is the hard case: the value is absent precisely because of what it would have been. An understanding of which regime you are in changes everything that follows — which tests to run, which imputer to trust, and how much uncertainty to carry forward into your downstream model.
What follows is a working guide to diagnosing and handling all three, with Python code you can adapt to real pipelines.
MCAR — the benign blank
"A sensor randomly drops a packet. A survey respondent accidentally skips a page. The phone dies mid-form. The value is gone for reasons entirely unrelated to the data itself."
MCAR is the statistician's dream and the practitioner's most overused assumption. When data is truly missing completely at random, the rows with missing values are an unbiased random sample of all rows. You can delete them without introducing systematic error. You can fill them with the column mean without distorting the distribution in any directional way. The danger is not in the technique — it is in the diagnosis. Many analysts assume MCAR because it is convenient, not because they have tested it.
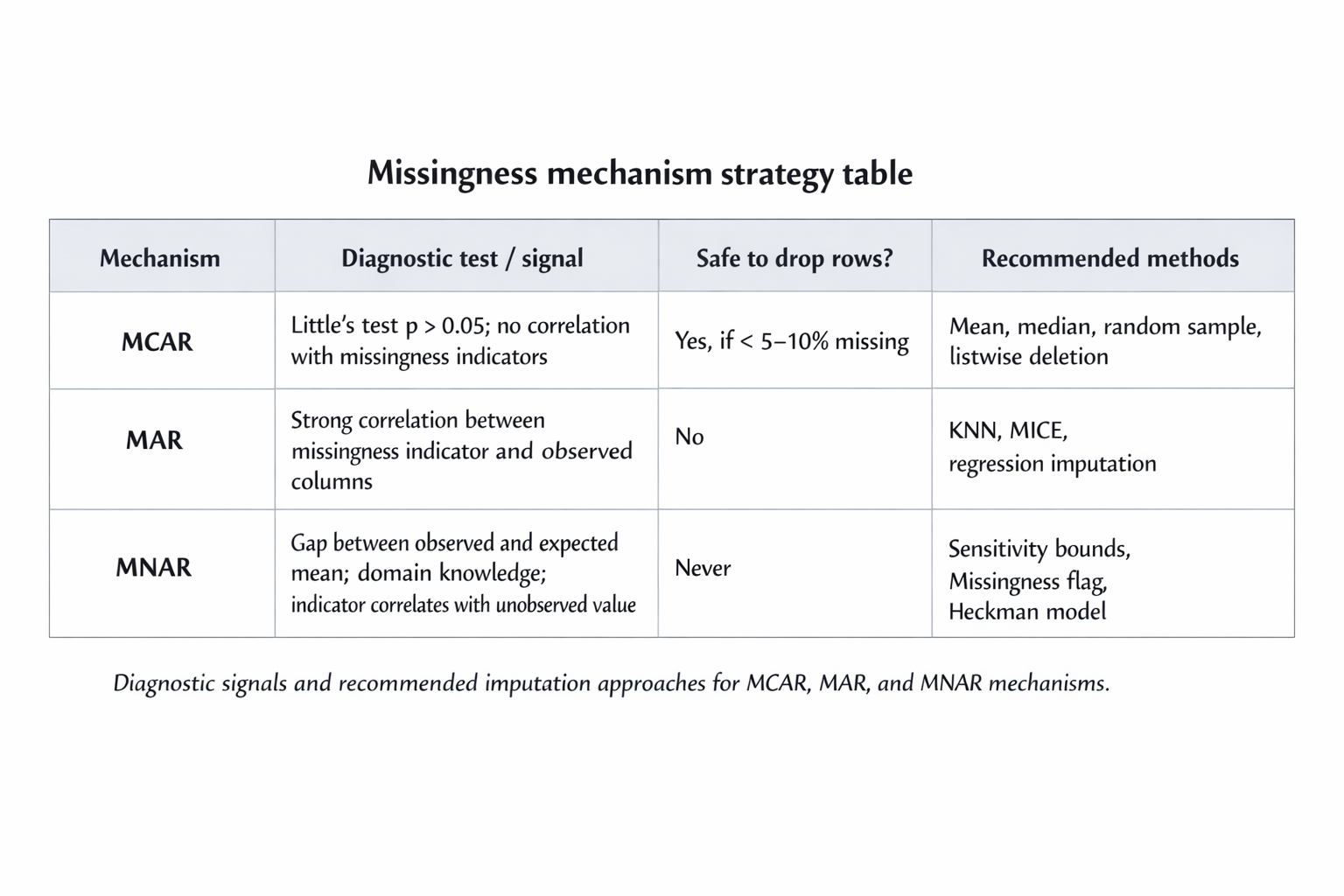
Little's MCAR test gives you a formal chi-squared statistic. A high p-value (conventionally above 0.05) provides evidence you cannot reject the MCAR hypothesis. It is not proof — it is a failure to disprove. Combine it with a visual missingness heatmap and an examination of whether missingness correlates with any observed column before you commit.
MCAR: Use cases
MCAR most commonly appears in IoT sensor streams with random dropout, large paper surveys with accidental skips, and automated data collection pipelines with transient network failures. It is far rarer in human-generated survey data on sensitive topics, where structured avoidance tends to dominate.
MCAR: Techniques
For MCAR, your choices are: mean or median imputation (fast, preserves column-level statistics), random sample imputation (preserves the full empirical distribution), or listwise deletion (removes the row entirely). The last option is only defensible when the percentage of missing rows is small — under five percent is a rough rule of thumb — and the dataset is large enough to absorb the loss..
MAR — the pattern hiding in plain sight
"Younger respondents skip the income question more often. Men underreport health scores. Patients with lower education levels leave clinical trial forms incomplete. The missing value has a pattern — but the pattern lives in columns you can see."
MAR is the workhorse case in real-world tabular data. The missingness is not random, but it is explainable by other observed variables. Once you condition on those variables, the remaining uncertainty is random. This means model-based imputation — which explicitly uses the observed columns to predict the missing ones — is both valid and effective. You are not guessing blindly; you are using information the data has already given you.
The classic diagnostic is to create a binary indicator column for each variable with missingness: income_missing = 1 where income is absent, 0 otherwise. Then check the correlation between that indicator and every other column in the dataset. A strong correlation tells you missingness is not random and points directly at which observed variables are driving the pattern. That is your MAR signature.
MAR: Use cases
MAR dominates in survey research (demographic characteristics predict non-response on sensitive questions), clinical datasets (sicker patients attend fewer follow-up visits, and severity is measured), credit scoring (employment status predicts missing income fields), and recommender systems (active users are observed more frequently than passive ones).
MAR: Techniques
KNN imputation uses the k nearest neighbours — measured by Euclidean distance across observed features — to fill each gap with a distance-weighted average of the neighbours' values. It is non-parametric and captures non-linear relationships, but scales poorly with dimensionality. MICE (Multiple Imputation by Chained Equations, implemented in scikit-learn as IterativeImputer) cycles through each column with missing values, fitting a regression model using all other columns as predictors and imputing from that model. It repeats this cycle until convergence. MICE is the gold standard for MAR data when you have the compute budget for it.
The KS test compares the imputed distribution to the observed distribution. A p-value above 0.05 is a rough sanity check that the imputer has not dramatically shifted the shape of the column. It is not definitive — the whole point is that missing values may have come from a different part of the distribution — but a very low p-value suggests the imputer is fabricating implausible values.
MNAR — the hole that knows its own shape
"High earners decline to report income. Patients who stop improving drop out of a clinical trial. People with the most debt skip the debt question. The value is absent because of what it would have been. The bias is structural."
MNAR is the case where no amount of clever imputation fully rescues you. The missingness is a function of the unobserved value itself, which means you cannot model it away using only the observed data — you would need to observe the very thing that is missing. This is a fundamentally different epistemological situation from MCAR or MAR, and it demands a different response: not confident imputation, but honest uncertainty quantification.
The practical tools are sensitivity analysis (what does my estimate look like under different assumptions about the missing values?), flagging missingness as an explicit binary feature in your model (which lets the model learn from the absence itself), and two-stage selection models like Heckman correction, which use the pattern of who is observed to partially correct for the selection bias. None of these eliminates the problem. All of them make the problem legible.
The telltale sign of MNAR is a gap between the observed mean and what you would expect the true mean to be from domain knowledge. If the average reported income in a survey is £42,000 but administrative records suggest the true average is £58,000, the people who chose not to report are disproportionately the high earners. That gap is the MNAR fingerprint.
MNAR: Use cases
MNAR appears wherever the act of reporting a value is itself informative: income and wealth surveys, mental health assessments, substance use questionnaires, attrition in longitudinal studies, online reviews (people who had neutral experiences rarely bother), and loan default data (applicants who would default are often the ones who do not complete applications).
MNAR: Techniques
Sensitivity analysis constructs a range of plausible estimates by filling missing values with optimistic, neutral, and pessimistic assumptions, then checking how much the downstream statistic (mean, model coefficient, risk estimate) varies across that range. If it barely moves, the missingness does not matter much. If it swings by 30%, you have a problem you need to communicate. The indicator method adds a variable_missing column to your feature matrix and fills the original with any fixed value (often the median). This lets tree-based models and neural networks learn that "this person did not answer" is itself a predictive signal. The Heckman selection model uses a two-stage approach: first model who gets observed, then use the inverse Mills ratio from that model as a bias-correction term in the outcome regression.
Choosing the right strategy
Explore more writing on topics that matter.